Hydractinia Genome Project Portal
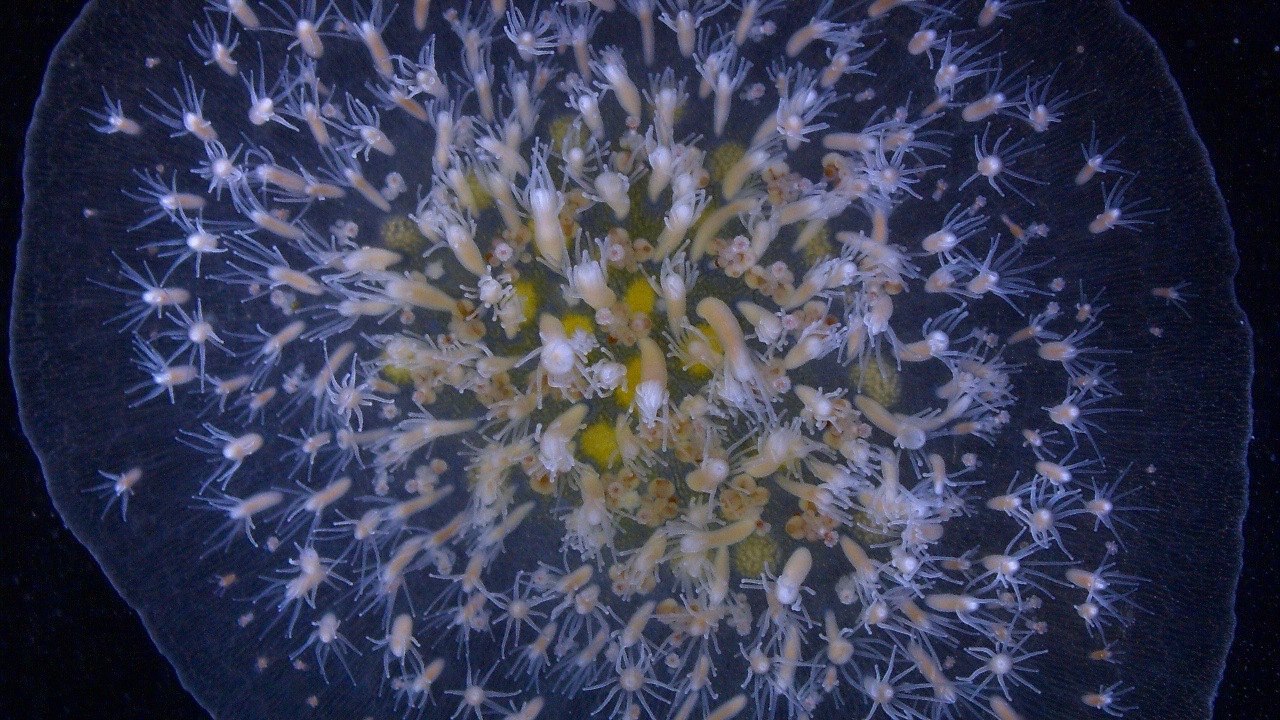
Marine colonial hydrozoans in the genus Hydractinia have a long history of biological study, dating back to August Weismann in the 1880s. From a phylogenetic standpoint, the cnidarians occupy an informative position on the animal tree as the sister group to the bilaterians. From a biological standpoint, members of the genus Hydractinia – hydrozoan cnidarians, or hydroids – are of particular interest, as they are amongst a very small number of organisms that can regenerate their entire body throughout their lifetime. This remarkable capacity for self-renewal is made possible due to migratory stem cells (called i-cells) that express genes whose bilaterian homologs are known to be involved in stem cell biology. Further, these colonial animals can distinguish between themselves and other members of their species purely based on cell-cell contact through a process called allorecognition.
Hydractinia has already proven to be amenable to experimental manipulation through techniques such as gene knockdown, CRISPR/Cas9 genome editing, and transgenesis, allowing researchers to address biological questions at the molecular level. Given their potential for studying key questions in regeneration, allorecognition, stem cell biology, and aging, we have sequenced the genomes of two Hydractinia species: H. echinata, which is native to the coastal waters of the northwestern Atlantic Ocean, and its sister species, H. symbiolongicarpus, which is native to the coastal waters of the northeastern Atlantic Ocean. The Hydractinia community has currently settled on using H. symbiolongicarpus as the main lab organism to perform most molecular-based studies.
This genome portal is intended to serve as a resource that provides genomic and transcriptomic data for researchers interested in Hydractinia genomics and molecular biology. These resources include browsers for the H. symbiolongicarpus and H. echinata genome assemblies and an interactive interface for interrogating the H. symbiolongicarpus single-cell gene expression atlas. We also provide gene pages with functional annotation data for every gene model for both species, including protein domain composition and orthology predictions.